By Kevin E. Noonan —
 On May 20th, Junior Party the University of California, Berkeley; the University of Vienna; and Emmanuelle Charpentier (collectively, "CVC") filed its Substantive Preliminary Motion No. 3 in Interference No. 106,127 (which names ToolGen as Senior Party), asking the Patent Trial and Appeal Board to add claims in ToolGen's U.S. Patent No. 10,851,380* to this interference, pursuant to 37 C.F.R. §§ 41.121(a)(1)(i) and 41.208(a)(2) and Standing Order ¶ 208.3.2.
On May 20th, Junior Party the University of California, Berkeley; the University of Vienna; and Emmanuelle Charpentier (collectively, "CVC") filed its Substantive Preliminary Motion No. 3 in Interference No. 106,127 (which names ToolGen as Senior Party), asking the Patent Trial and Appeal Board to add claims in ToolGen's U.S. Patent No. 10,851,380* to this interference, pursuant to 37 C.F.R. §§ 41.121(a)(1)(i) and 41.208(a)(2) and Standing Order ¶ 208.3.2.
Relevant to CVC's motion is the portion of the Count identical to Claim 85 of ToolGen's priority application, U.S. provisional application No. 14/685,510:
An isolated mammalian cell comprising a Type II Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas system for site specific, cleavage of a double-stranded target nucleic acid sequence in the isolated mammalian cell, wherein the CRISPR/Cas system comprises:
a) a nucleic acid encoding a Cas9 polypeptide, wherein the Cas9 polypeptide comprises a nuclear localization signal and wherein said nucleic acid is codon-optimized for expression in mammalian cells, and
b) a chimeric guide RNA comprising a CRISPR RNA (crRNA) portion and a transactivating crRNA (tracrRNA) portion,
wherein the target nucleic acid sequence comprises a first strand having a region complementary to the crRNA portion of the chimeric guide RNA and a second strand having a trinucleotide protospacer adjacent motif (PAM), and
wherein the Cas9 polypeptide and the chimeric guide RNA form a Cas9/RNA complex in the isolated mammalian cell and mediate double stranded cleavage at the target sequence.
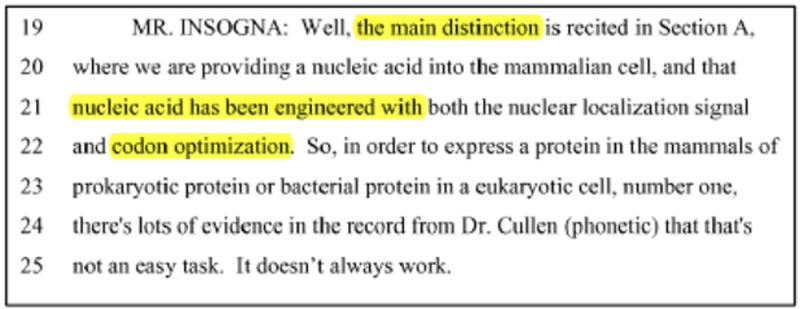
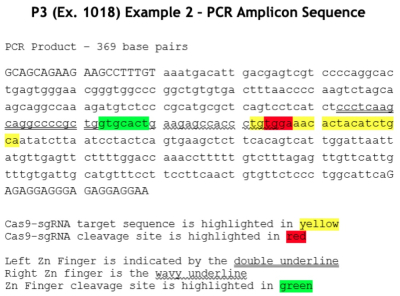
CVC argues that the only difference between the language of the Count and the claims in the '380 patent is that those claims require the addition of two guanine residues ("GG") positioned before the crRNA portion of the sgRNA sequence. CVC argues that these species of sgRNA (the fusion of crRNA and tracrRNA) recited in the '380 patent claims are a consequence of using the T7 phage RNA polymerase to produce sgRNA and that in vitro RNA production using T7 RNAP promoters was well-known in the art ("for decades"; emphasis in brief) at the priority date of the '380 patent; these arguments are supported by testimony from CVC's expert, Dr. Scott Bailey.
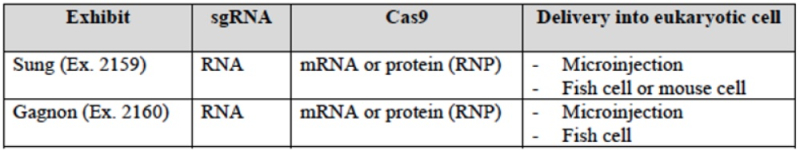
This method of producing sgRNA and relevant prior art disclosing the use of T7 RNAP and promoters recognized by the polymerase are set forth in the brief as follows:
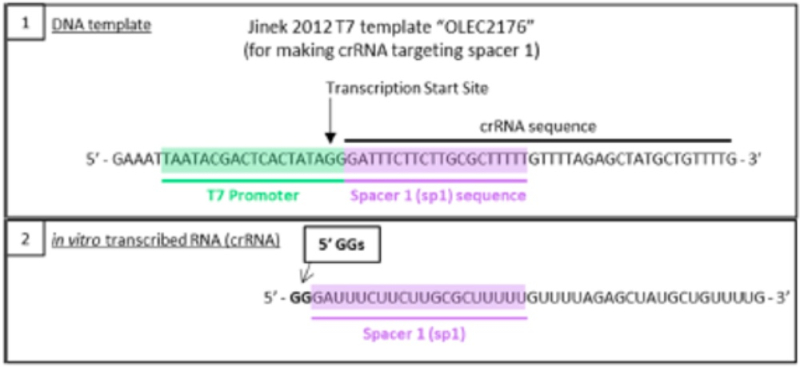
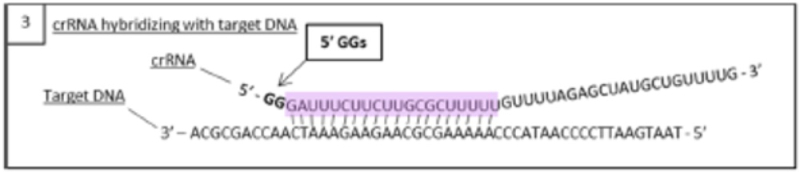
wherein the diagrams follow the source of the characteristic "extra" GG residues at the 5' end of crRNA and sgRNA produced thereby. CVC also asserts a second reference, the Deltcheva reference, to illustrate the extent to which T7-catalyzed in vitro RNA production using a T7 promoter was well-known:
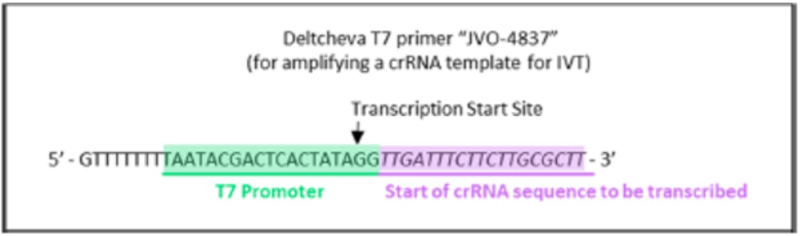
CVC argues that the distinction of including two guanine residues in crRNA and sgRNA comprised thereof is not enough to distinguish the claims of the '380 patent from the Count in this interference (to which CVC argues these claims correspond) because "including a 5'-GG would have been obvious over Count 1 in view of [CVC's] Jinek 2012" reference as illustrated above, which reliance is permitted under Desjardins v Wax, Interference No. 105,915, Paper 127, 17-20 (P.T.A.B. Jan. 21, 2014). Regarding motivation to combine the teachings of the Jinek reference in this regard with the more general teachings of producing an sgRNA for eukaryotic CRISPR, CVC argues that such motivation is supported by the method's "low cost, efficiency, and accuracy," and because "Jinek 2012 had already used the method to generate RNAs that effectively cleave eukaryotic DNA sequences (e.g., GFP) in CRISPR-Cas9 systems (i.e., ToolGen knew the method would be successful in eukaryotic CRISPR). This success also would have provided the requisite reasonable expectation of success to complete a prima facie case of obviousness and hence for the '380 claims to properly be determined to correspond to the Count in this interference.
The brief sets forth in detail comparison between the Count and claims of the '380 patent to set forth CVC's argument that those claims would have been obvious over the Count (considered as part of the prior art for this analysis; see N.V. Nutricia v. Mass. Inst. of Tech., Interference No. 106,096, Paper 80, 4 (P.T.A.B. Mar. 29, 2019)) in combination, inter alia, the Jinek 2012 reference (or any other reference teaching the use of the T7 promoter/ T7 RNA polymerase system for producing RNA including sgRNA in vitro. CVC characterizes its argument by stating that those claims "the predictable use of prior art elements according to their established functions," citing KSR Intern. Co. v. Teleflex Inc., 127 S.Ct. 1727, 1731 (2007); these arguments are supported by a detailed claim chart set out as Appendix 3 to CVC's brief (providing the comparison with the portion of the Count taken from ToolGen's application). (Appendix 4 contains a similar claim chart from the portion of the Count taken from claim 156 of the CVC's U.S. Application No. 15/981,807.)
Turning to the basis for CVC's assertion of there being a reasonable expectation of success, CVC argues that:
Expectation of success in eukaryotes would not be in doubt because "whether a CRISPR-Cas9 system would have been expected to work in a eukaryotic cell . . . is assumed under the framework of 37 C.F.R. 41.207(b)(2), wherein Count 1 is presumed to be prior art to the . . . claims," citing The Univ. of Calif. v. The Broad Inst., Inc., Interference No. 106,115, Paper 877, 66 (P.T.A.B. Sept. 10, 2020); 37 C.F.R. 41.207(b)(2).
Having set forth the basis for the Board to find claim 1 obvious, the brief then sets forth CVC's obviousness analysis regarding claims 2-10 as having been obvious (and the brief is supplemented with Appendix showing those sites).
The brief completes its obviousness assessment by asserting that there was no objective indicia that would contradict the prima facia case of obviousness supported by the facts and argument set forth above. These include specifically the absence of any evidence of unexpected results (which ToolGen argued during prosecution of the '380 patent) in view of (for the purposes of this interference) ToolGen's half of the Count, which differs from the '380 claims only by the addition of the GG residues at the 5' end of the sgRNA as a consequence of producing that RNA species in vitro using T7 RNA polymerase and its cognate promoter. CVC arrives at this conclusion because the claim corresponding to ToolGen's half of the interference Count (claim 85 of ToolGen's involved '510 application) was prosecuted to allowance, inter alia, by ToolGen's argument for unexpected results; as CVC asserts "there can be no unexpectedly superior results when the closest prior art (i.e., Count 1) has the same results as the claimed method," citing Millennium Pharmaceuticals, Inc. v. Sandoz, Inc., 862 F.3d 1356, 1368 (Fed. Cir. 2017) (emphasis in brief). In addition, CVC argues that here the prima facie case is sufficiently strong so as not to be rebutted by unexpected results, citing Ohio Willow Wood Co. v. Alps South, LLC, 735 F.3d 1333, 1344 (Fed. Cir. 2013) ("[W]here a claimed invention represents no more than the predictable use of prior art elements according to established functions, as here, evidence of secondary indicia are frequently deemed inadequate to establish nonobviousness."). Finally, merely discovering the effect of adding the GG residues at the 5' end of the sgRNA is not sufficient to overcome a prima facie obviousness case, based on Federal Circuit precedent that a "'previously-unknown, yet inherent, food-effect property' did not make the claims patentable because 'merely discovering and claiming a new benefit of an old process cannot render the process again patentable,'" citing In re Huai-Hung Kao, 639 F.3d 1057 (Fed. Cir. 2011).
Finally, CVC's brief concludes with its argument that there is an interference-in-fact between CVC's involved claims in this interference and the '380 patent claims (illustrating the principle that once bitten is twice shy). CVC argues here that the "two-way" test for interfering subject matter is met between at least one claim of the '380 patent and at least one of CVC's involved claims. Specifically, the brief calls out claim 1 of the '380 patent and claim 156 of CVC's involved application No. 15/981,807:
156. A eukaryotic cell comprising a target DNA molecule and an engineered and/or non-naturally occurring Type II Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-CRISPR associated (Cas) (CRISPR-Cas) system comprising
a) a Cas9 protein, or a nucleic acid comprising a nucleotide sequence encoding said Cas9 protein; and
b) a single molecule DNA-targeting RNA, or a nucleic acid comprising a nucleotide sequence encoding said single molecule DNA-targeting RNA;
wherein the single molecule DNA-targeting RNA comprises:
i) a targeter-RNA that is capable of hybridizing with a target sequence in the target DNA molecule, and
ii) an activator-RNA that is capable of hybridizing with the targeter-RNA to form a double-stranded RNA duplex of a protein-binding segment,
wherein the activator-RNA and the targeter-RNA are covalently linked to one another with intervening nucleotides; and
wherein the single molecule DNA-targeting RNA is capable of forming a complex with the Cas9 protein, thereby targeting the Cas9 protein to the target DNA molecule,
whereby said system is capable of cleaving or editing the target DNA molecule or modulating transcription of at least one gene encoded by the target DNA molecule.
CVC identifies as "the only relevant differences" between this claim and claim 1 of ToolGen's '380 patent:
• codon optimization of the Cas9 polynucleotide,
• the NLS-tag on Cas9, and
• including a 5'-GG on the guide RNA.
Noting that claim 185 of CVC's '807 application recites use of the NLS tag, CVC argues that the skilled worker would have had a reasoned basis for including all of these features into a CRISPR-Cas9 system, under the rubric that a "combination of familiar elements according to known methods is likely to be obvious when it does no more than yield predictable results," citing KSR Intern. Co. v. Teleflex Inc., 127 S.Ct. 1727, 1731 (2007). And, CVC maintains, the skilled worker would have had a reasonable expectation of success in achieving operative embodiments of CRISPR thereby, satisfying one prong of the two-way test.
With regard to the other prong, CVC argues that claim 1 of ToolGen's '380 patent teaches all the elements of CVC's claim 156 in the order in which it is claimed, which is enough to satisfy the test for anticipation.
Accordingly, CVC concludes, the Board should designate the claims of the '380 to correspond to Count 1, adding that "[t]here are no adequate alternative remedies because leaving the '380 patent out of this proceeding risks the inefficiency and expense of another interference proceeding involving CVC and ToolGen."
* '830 Patent claims CVC asserts correspond to the Count in the '127 Interference:
1. A method of introducing a site-specific, double-stranded break at a target nucleic acid sequence in a eukaryotic cell, the method comprising introducing into the eukaryotic cell a Type II Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas system, wherein the CRISPR/Cas system comprises: a) a nucleic acid encoding a Cas9 polypeptide comprising a nuclear localization signal, wherein the nucleic acid is codon-optimized for expression in eukaryotic cells, and b) a guide RNA that hybridizes to the target nucleic acid, wherein the guide RNA is a chimeric guide RNA comprising a CRISPR RNA (crRNA) portion fused to a trans activating crRNA (tracrRNA) portion, wherein the guide RNA comprises two guanines at its 5' end, and there are no additional nucleic acid residues between the two guanines at the 5' end and the crRNA portion of the guide RNA; whereby a site-specific, double stranded break at the target nucleic acid sequence is introduced.
2. The method of claim 1, wherein the nuclear localization signal is located at the C terminus of the Cas9 polypeptide.
3. The method of claim 1, wherein the eukaryotic cell is a mammalian cell.
4. The method of claim 3, wherein the mammalian cell is a human cell.
5. The method of claim 1, wherein the nucleic acid encoding the Cas 9 polypeptide is codon-optimized for expression in mammalian cells.
6. The method of claim 1, wherein the target nucleic acid sequence is a genomic sequence located at its endogenous site in the genome of the eukaryotic cell.
7. The method of claim 1, wherein the nucleic acid encoding the Cas9 polypeptide is a vector.
8. The method of claim 1, wherein the Cas9 polypeptide is a Streptococcus pyogenes Cas9 polypeptide.
9. The method of claim 1, wherein the nucleic acid encoding the Cas9 polypeptide is introduced into the eukaryotic cell before introducing the guide RNA into the eukaryotic cell.
10. The method of claim 1, wherein the Cas9 polypeptide is a Streptococcus Cas9 polypeptide.
 IPWatchdog and Foresight and will be offering a webinar entitled "We Did It Our Way: Women IP Trailblazers Share Their Incredible Journeys" on July 13, 2021 at 2:00 pm (ET). Gene Quinn of IPWatchdog, Inc. will moderate a panel consisting of Angela Grayson, Founder and Principal Member, PRECIPICE; Mary Jutten, CEO Traklight, Inc and Juetten Law, P.C.; Efrat Kasznik, Founder and Founder & President, Foresight Valuation Group, LLC; and Renée C. Quinn, Chief Operating Officer & Chief Financial Officer, IPWatchdog, Inc. The panel will address questions such as what a "typical" career in IP entails and how women are faring as IP professionals.
IPWatchdog and Foresight and will be offering a webinar entitled "We Did It Our Way: Women IP Trailblazers Share Their Incredible Journeys" on July 13, 2021 at 2:00 pm (ET). Gene Quinn of IPWatchdog, Inc. will moderate a panel consisting of Angela Grayson, Founder and Principal Member, PRECIPICE; Mary Jutten, CEO Traklight, Inc and Juetten Law, P.C.; Efrat Kasznik, Founder and Founder & President, Foresight Valuation Group, LLC; and Renée C. Quinn, Chief Operating Officer & Chief Financial Officer, IPWatchdog, Inc. The panel will address questions such as what a "typical" career in IP entails and how women are faring as IP professionals.