By Kevin E. Noonan —
 Judge Giles Sutherland Rich's most famous aphorism in patent law is "the name of the game is the claim."* This rubric is important to keep in mind when considering the Patent Trial and Appeal Board's decision on motions issued September 10th in Interference No. 106,115 (see "PTAB Decides Parties' Motions in CRISPR Interference") between Senior Party The Broad Institute, Harvard University, and the Massachusetts Institute of Technology (collectively, "Broad") and Junior Party the University of California/Berkeley, the University of Vienna, and Emmanuelle Charpentier (collectively, "CVC").
Judge Giles Sutherland Rich's most famous aphorism in patent law is "the name of the game is the claim."* This rubric is important to keep in mind when considering the Patent Trial and Appeal Board's decision on motions issued September 10th in Interference No. 106,115 (see "PTAB Decides Parties' Motions in CRISPR Interference") between Senior Party The Broad Institute, Harvard University, and the Massachusetts Institute of Technology (collectively, "Broad") and Junior Party the University of California/Berkeley, the University of Vienna, and Emmanuelle Charpentier (collectively, "CVC").
The focus of the claim construction issue was the meaning of the term "guide RNA," specifically whether it was a generic term encompassing dual-species or single-species ("fused") RNA components of the CRISPR-Cas9 complex, as illustrated in the Decision (taken from the Jinek 2012 reference; Jinek et al., 2012, "A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity," Science 337: 816–21):
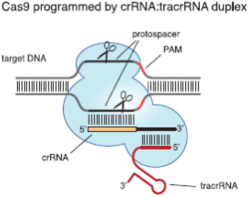

In its Motions Nos. 2 and 3, Broad asserted this distinction, relying on portions of the specifications of several of its patents-in-interference. While the term "crRNA" is recited in these Broad specifications, the involved claims recite "guide RNA," "chimeric RNA," and "guide sequence," and as the Decision states, the parties disputed the meaning of this term. Broad noted that "the majority" of their involved claims recite "guide RNA" and that term was not limited to single- or dual-(RNA) molecule embodiments. CVC, for its part, argued that the term should be limited to single-(RNA) molecule species for all Broad's claims-in-interference.
The Board used the "broadest reasonable interpretation" test (that standard was not changed when the Patent and Trademark Office adopted the Phillips test for post-grant review proceedings; see "PTAB Adopts Litigation Standards for Claim Construction in AIA Proceedings"), noting that in applying that standard "the Board's construction cannot be divorced from the specification and the record evidence, and must be consistent with the one that those skilled in the art would reach," citing Microsoft Corp. v. Proxycom, Inc., 789 F.3d 1292, 1298 (Fed. Cir. 2015).
Broad's argument in favor of construing "guide RNA" as a generic term encompassing both single- and dual-molecule embodiments was based on claims that distinguished between "guide RNA" and "fused guide RNA." Under the principle of claim differentiation, Broad argued that a claim reciting "fused guide RNA" that was dependent on a claim reciting "guide RNA" indicates that the latter term is broader than "fused" guide RNA (citing claim 18, dependent on claim 15, of Broad's Patent No. 8,697,359 in support of this argument). Similarly, Broad cited claim 3 of its Patent No. 8,993,233 that recites a limitation wherein the guide RNA comprises a tracrRNA sequence on a different vector than the nucleic acid encoding the Cas9 protein.
CVC in opposition countered that claim differentiation is not "a rigid rule" and the segregation of guide RNA and Cas9-encoding RNA in the '233 patent did not mandate either guide RNA configuration.
The Board stated that while it agrees with Broad that some of its claims suggest that the term "guide RNA" is generic, in other instances the claims are amenable to a contrary construction of the term. Accordingly, the Decision went on to consider other evidence to see if it rebutted the presumption that these claims had raised.
That evidence included support for Broad's argument that there is no "clear disavowal of scope" not to include both species of guide RNAs in the claims. Broad contended that the phrase is a "term of art" having a plain meaning that encompasses both single- and dual-RNA species. Broad relied on the Jinek reference for support, citing this figure:
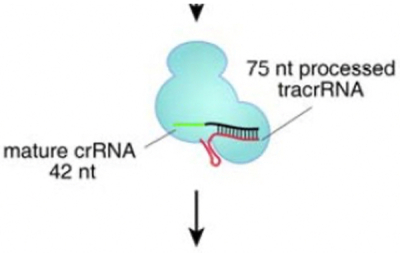 accompanied by the caption "In this ternary complex, the dual tracrRNA:crRNA structure acts as guide RNA that directs the endonuclease Cas9 to the cognate target DNA" (emphasis in opinion). Broad further supported this argument with expert testimony (consistent with Broad's interpretation of the figure) that:
accompanied by the caption "In this ternary complex, the dual tracrRNA:crRNA structure acts as guide RNA that directs the endonuclease Cas9 to the cognate target DNA" (emphasis in opinion). Broad further supported this argument with expert testimony (consistent with Broad's interpretation of the figure) that:
The "ternary complex" refers to the three part complex that consists of (1) Cas9, (2) a mature crRNA molecule and (3) a tracrRNA molecule. This is significant because "the dual tracrRNA:crRNA structure" makes up two parts of the three part complex. Otherwise, Jinek 2012 would not have referred to the Cas9:RNA complex as a "ternary complex," but as a binary complex. Thus, the "guide RNA" in that sentence references the dual-guide RNA consisting of separate strands of tracrRNA and crRNA.
The Board found other evidence from the Broad to be "less persuasive." This included expert testimony evaluating CVC's specification in U.S. Serial No. 15/947,680 as reciting "[t]he term 'DNA-targeting RNA' or 'gRNA' is inclusive, referring both to double-molecule DNA-targeting RNAs and to single-molecule DNA-targeting RNAs (i.e., sgRNAs)," which the Board professed was not conclusive with regard to an interpretation that the meaning of "gRNA" meant "guide RNA." Similarly, Broad's expert testified that CVC inventors used the term "guide RNA" inclusively to mean "all crRNA:tracrRNA complexes, whether present as a single or a double-molecule" in a scientific paper (Sternberg et al., 2014, "DNA interrogation by the CRISPR RNA-guided endonuclease Cas9," Nature 507: 62), but the Board stated that "without an explanation of how these specific instances of the terms would have been understood by those in the art at the time, we are not certain they demonstrate uses of 'guide RNA' as a generic term." The same expert also cited CVC witness testimony from earlier Interference No. 105,048 that "guide DNA targeting RNA" includes either single- or dual-guide species, but the Board noted that this term is not the same as the "guide RNA" term as used in Broad's motions in this interference. And the Board rejected out of hand similar statements by CVC counsel in the '048 interference, because counsel "is not one of skill in the art and we are not persuaded that his use of technical terms indicates anything about how they would have been understood by those in the art at the time."
The Board then turned to Broad's assertion of extrinsic evidence, including several contemporaneous scientific journal articles and references, all of which (being published prior to 2011) the Board found irrelevant. In summary the Board states that:
We are not persuaded from the extrinsic evidence cited by Broad that the term "guide RNA" was well known in the art to mean either a single or a dual RNA molecule configuration. In some publications cited by Broad, such as Jinek 2012, the term "guide RNA" is used to refer to a dual molecule RNA configuration. But in other examples, such as CVC's '680 application and Drs. Geider and Carroll's declarations in the prior '048 interference, the specific term "guide RNA" was not actually used. In yet other references, such as Bhaya, Horvath, Rand, and Tolia, the term is used, but not for a complex of the crRNA and tracrRNA. This evidence does not persuade us that the term "guide RNA" had a plain meaning in the art, which "indisputably" included both single- and dual- molecule RNA configurations, as Broad argues.
From this determination that the term "guide RNA" was not understood to be generic at the time of Broad's filings, the Board states that it is not convinced that "the specification must provide a clear intent to exclude a dual-molecule RNA configuration from the term," citing Trs. of Columbia Univ. v. Symantec Corp., 811 F.3d 1359, 1363 (Fed. Cir. 2016), for an explication of the requirements under Phillips v. AWH Corp., 415 F.3d 1303, 1320 (Fed. Cir. 2005 (en banc), in this regard. Broad cited a portion of its specification in support of its argument:
In aspects of the invention the terms "chimeric RNA", "chimeric guide RNA", "guide RNA", "single guide RNA" and "synthetic guide RNA" are used interchangeably and refer to the polynucleotide sequence comprising the guide sequence, the tracr sequence and the tracr mate sequence. The term "guide sequence" refers to the about 20 bp sequence within the guide RNA that specifies the target site and may be used interchangeably with the terms "guide" or "spacer". The term "tracr mate sequence" may also be used interchangeably with the term "direct repeat(s)". An exemplary CRISPR-Cas system is illustrated in FIG. 1.
Broad argued this portion of the specification referenced some (but not all) aspects of the invention to encompass single RNA guide RNA species. Broad also argued that "used interchangeably" does not mean that the listed terms are themselves the same molecules. CVC disputed these characterizations, arguing that "the specification specifically states that the terms 'guide RNA,' 'chimeric RNA,' 'chimeric guide RNA,' and 'single guide RNA' all 'refer to the polynucleotide sequence comprising the guide sequence, the tracr sequence and the tracr mate sequence.'" And CVC further argued that "this portion of the specification defines "guide RNA" as a singular polynucleotide sequence comprising a guide sequence, a tracr sequence, and a tracr mate sequence and corresponding to the fused crRNA and the tracrRNA."
These arguments were persuasive to the Board, which stated that "this paragraph of the Broad specification indicates that 'chimeric RNA,' 'chimeric guide RNA,' 'single guide RNA,' as well as 'guide RNA' include these three components."
The Decision further explicated other portions of the various Broad specifications, to the same effect: the Board is persuaded by CVC's arguments (or not persuaded by Broad's arguments) regarding the status of the term "guide RNA" as a generic term encompassing both single- and dual-RNA species.
The Board offers this conclusion regarding its construction of this sole term:
Our review of the parties' arguments leads us to the conclusion that Broad's use of the term "guide RNA" in its involved claims is not a generic term, but is limited to a single-molecule RNA configuration of the guide sequence and tracrmate, which together make the crRNA, and the tracrRNA sequences. Although some dependent claims, such as claim 18 of the '359 patent, might indicate by claim differentiation that the term "guide RNA" is generic, that presumption is overcome by Broad's specification. The specification of Broad's involved patents, specifically the sentence providing that "guide RNA" and other terms "refer to the polynucleotide sequence comprising the guide sequence, the tracr sequence and the tracr mate sequence" ('359 patent, Ex. 3011, 12:6–10), limits the interpretation of the term. Broad fails to direct us to other uses of the term "guide RNA" in the specification that indicate a dual-molecule RNA configuration and we are not persuaded that the term was so clearly understood the art to be a generic term that only a clear disavowal in the specification would define it to mean a single-molecule RNA configuration. Thus, we are persuaded that the broadest reasonable interpretation of Broad claim term "guide RNA" encompasses only a single-molecule RNA configuration.
The consequences of this claim construction will be the subject of future posts.
*And then there is this ditty sung to the tune of "Camelot":
A law was made 200 years ago here
Grant patents, help promote inventive thought
Today the system's thriving and our credo
Is claim-a-lot
We push the envelope,
Expand the boundaries
Create a circle from a tiny dot
Our product's forged with words
and not in foundries
We claim a lot
Claim-a-lot (claim-a-lot)
I know it sounds a bit bizarre
Lord, we claim-a-lot (oh yes, we claim-a-lot)
Stretch out those claims so far
Though prior art may set some limitations
Restricts our flights of fancy, clever thought
Our efforts, not for naught
Results, so boldly wrought
Construct our patent juggernauts
By claiming quite a lot.
Kramer, Levin, Naftalis & Frankel LLP, Claim-a-Lot, in Pamphlet for N.Y. Intellectual Property Law Association 78th Annual Dinner (Mar. 24, 2000).
 LexisNexis IP and IPWatchdog and will be offering a webinar entitled "Myths of Litigated Patents" on October 15, 2020 at 12:00 pm (ET). Dr. Sean Tu, Professor, WVU College of Law; Megan McLoughlin of LexisNexis® IP; and Gene Quinn of IPWatchdog will discuss whether patent examiners who issue litigated patents have common characteristics? While intuition would argue that those examiners who issue the most patents (approximately one patent every three business days) would exhibit a higher litigation rate, surprisingly, two studies by Professor Sean Tu suggest that this is wrong. The panel will address the following issues:
LexisNexis IP and IPWatchdog and will be offering a webinar entitled "Myths of Litigated Patents" on October 15, 2020 at 12:00 pm (ET). Dr. Sean Tu, Professor, WVU College of Law; Megan McLoughlin of LexisNexis® IP; and Gene Quinn of IPWatchdog will discuss whether patent examiners who issue litigated patents have common characteristics? While intuition would argue that those examiners who issue the most patents (approximately one patent every three business days) would exhibit a higher litigation rate, surprisingly, two studies by Professor Sean Tu suggest that this is wrong. The panel will address the following issues: